| 549 | | To try this we need a more complex topology. The program below creates a topology with {{{nrouters}}} completely connected routers, each of which has {{{nleaves}}} leaves, except for one central router that has none. |
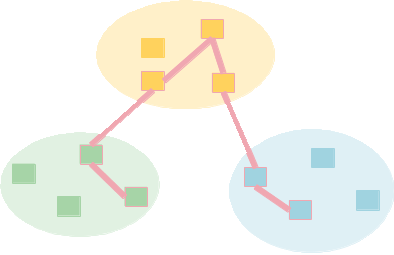
| | 549 | To try this we need a more complex topology. The program below creates a topology with {{{nrouters}}} completely connected routers, each of which has {{{nleaves}}} leaves, except for one central router that has none. Here is an image of the topology when {{{nrouters}}}=4 and {{{nleaves}}}=4: |
| | 550 | |
| | 551 | [[Image(topo.png)]] |
| | 552 | |
| | 553 | We leave one router without leaves for aecthetics. |
| | 554 | |
| | 555 | Here is the program that generates that topology: |
| | 556 | |
| | 557 | {{{ |
| | 558 | #!/usr/bin/env python |
| | 559 | |
| | 560 | import sys |
| | 561 | from deter import topdl |
| | 562 | |
| | 563 | # Parse the user-supplied routers and leaves arguments |
| | 564 | try: |
| | 565 | if len(sys.argv) > 1: arg = sys.argv[1] |
| | 566 | else: arg = 4 |
| | 567 | nrouters = int(arg) |
| | 568 | |
| | 569 | if len(sys.argv) > 2: arg = sys.argv[2] |
| | 570 | else: arg = 3 |
| | 571 | nleaves = int(arg) |
| | 572 | |
| | 573 | except ValueError, e: |
| | 574 | print >>sys.stderr, 'Usage %s routers leaves' % sys.argv[0] |
| | 575 | sys.exit('Cannot convert %s to an int' % arg) |
| | 576 | |
| | 577 | elems = [] # Elements in the topology |
| | 578 | subs = [] # Substrates in the topology |
| | 579 | |
| | 580 | # Make nrouters routers |
| | 581 | r = [] |
| | 582 | for i in range(0,nrouters): |
| | 583 | r.append(topdl.Computer(name='r-%d' % i)) |
| | 584 | |
| | 585 | elems.extend(r) |
| | 586 | # Interconnect them |
| | 587 | n = 0 |
| | 588 | for i in range(0,nrouters): |
| | 589 | for j in range(i+1, nrouters): |
| | 590 | # Connect r[i] to r[j] |
| | 591 | # Pick a substrate name |
| | 592 | link_name = 'rlink%i' % n |
| | 593 | # Add interfaces to the two routers |
| | 594 | r[i].interface.append(topdl.Interface(substrate=[link_name])) |
| | 595 | r[j].interface.append(topdl.Interface(substrate=[link_name])) |
| | 596 | # Put a substrate into the substrate list with the name we picked |
| | 597 | subs.append(topdl.Substrate(name=link_name)) |
| | 598 | n += 1 |
| | 599 | |
| | 600 | # Put nleaves leaves on each router, except the center one |
| | 601 | for i in range(0, (nrouters-1) * nleaves): |
| | 602 | # Pick a link name |
| | 603 | link_name = 'llink%i' % i |
| | 604 | |
| | 605 | # add a computer with an interface on the new substrate to the element list |
| | 606 | elems.append(topdl.Computer(name='leaf%d' % i, interface=[ |
| | 607 | topdl.Interface(substrate=[link_name])])) |
| | 608 | |
| | 609 | # Add an interface to a router on the new substrate |
| | 610 | r[i/nleaves].interface.append(topdl.Interface(substrate=[link_name])) |
| | 611 | |
| | 612 | # Put a substrate into the substrate list with the name we picked |
| | 613 | subs.append(topdl.Substrate(name=link_name)) |
| | 614 | |
| | 615 | # Make the topology and print it |
| | 616 | top = topdl.Topology(elements=elems, substrates=subs) |
| | 617 | print topdl.topology_to_xml(top, top='experiment') |
| | 618 | |
| | 619 | }}} |
| | 620 | |
| | 621 | The code takes a few more shortcuts in construction, e.g., most elements and substrates are appended directly to the lists rather than being assigned to variables, but is not fundamentally different from the start topology generator. |